Soluble Guanylate Cyclase
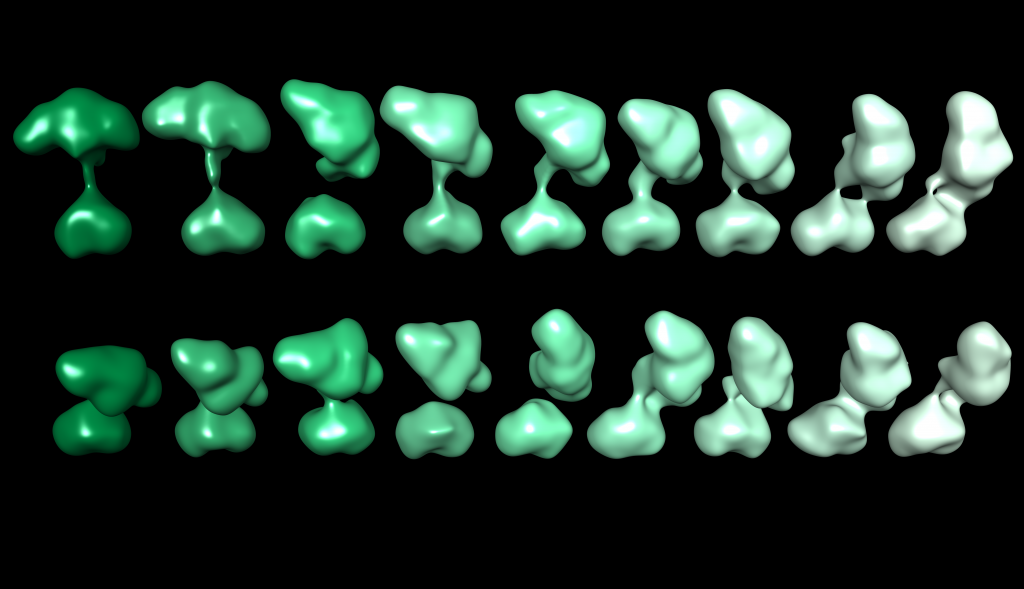 Soluble guanylate cyclase or “sGC” is a small protein that serves as an important part of the nitric oxide (NO) signaling pathway in mammals. Disruptions in this pathway lead to a large number of diseases including heart disease and stroke — two of the leading causes of death in America today. Several drugs targeting the activity of sGC are currently being investigated in clinical trials. In order to better understand sGC and the way these drugs work, we study the structure using a high-power electron microscope. Many images of the protein at a magnification of 80,000x were acquired using a method called “random conical tilt.” In this method, we image each particle twice: once at no tilt, and once at a tilted angle. This allows us to use computational analysis to sort and align these images to reconstruct a three-dimensional structure. We found that sGC is very dynamic and constantly moving. The image here shows 18 ‘snapshots’ highlighting the range of motion available to sGC.
Soluble guanylate cyclase or “sGC” is a small protein that serves as an important part of the nitric oxide (NO) signaling pathway in mammals. Disruptions in this pathway lead to a large number of diseases including heart disease and stroke — two of the leading causes of death in America today. Several drugs targeting the activity of sGC are currently being investigated in clinical trials. In order to better understand sGC and the way these drugs work, we study the structure using a high-power electron microscope. Many images of the protein at a magnification of 80,000x were acquired using a method called “random conical tilt.” In this method, we image each particle twice: once at no tilt, and once at a tilted angle. This allows us to use computational analysis to sort and align these images to reconstruct a three-dimensional structure. We found that sGC is very dynamic and constantly moving. The image here shows 18 ‘snapshots’ highlighting the range of motion available to sGC.
The image was rendered by Melody Campbell (TSRI).
Work that led to the 3D map was published in:
- Campbell MG, Underbakke ES, Potter CS, Carragher B, Marletta MA. Single-particle EM reveals the higher-order domain architecture of soluble guanylate cyclase. Proc Natl Acad Sci U S A. 2014;111(8):2960-5. PubMed
